|
Yu Zhang, Wei Wu, Russell T. Toll, Sharon Naparstek, Adi Maron-Katz, Mallissa Watts, Joseph Gordon, Jisoo Jeong, Laura Astolfi, Emmanuel Shpigel, Parker Longwell, Kamron Sarhadi, Dawlat EI-Said, Yuanqing Li, Crystal Cooper, Cherise Chin-Fatt, Martjn Arns, Madhukar H. Trivedi, Charles R. Marmar, Amit Etkin
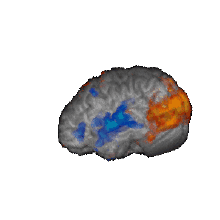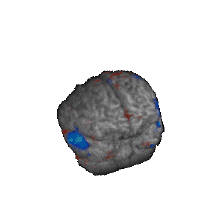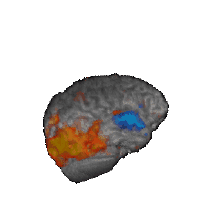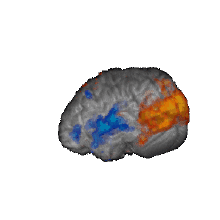
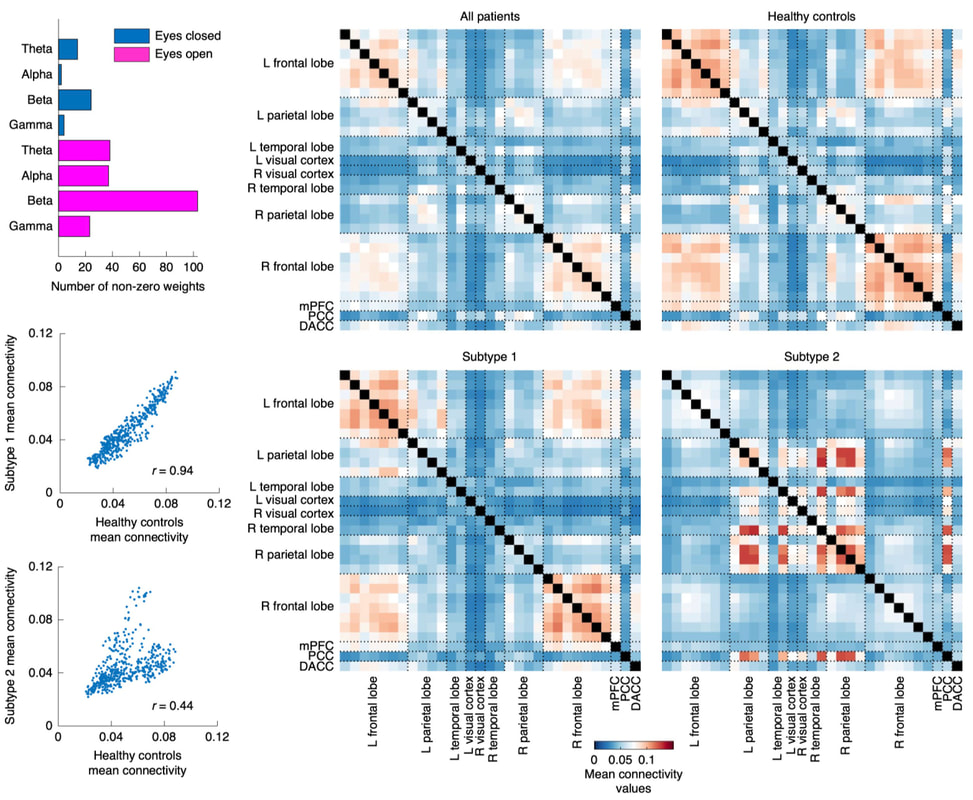
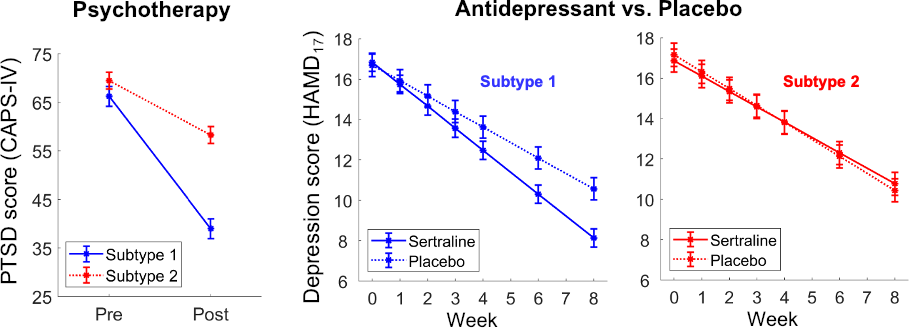
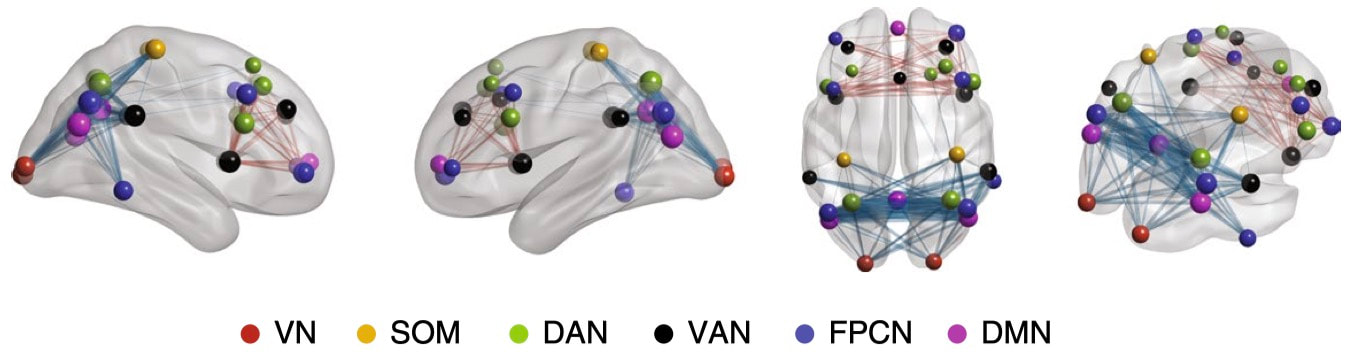
Nature Biomedical Engineering, 2021 Motivation: The understanding and treatment of psychiatric disorders are known to be neurobiologically and clinically heterogeneous but could benefit from the data-driven identification of disease subtypes. Solution: We developed a data-driven subtyping framework based on unsupervised sparse clustering to accomplished simultaneous feature selection and sample clustering on the power-envelope-based connectivity of signals reconstructed from high-density resting-state EEG. Extensive validation and replication were performed on four independent datasets involving two different psychiatric disorders, including PTSD and MDD. Results: We identified the disease subtypes and showed that the subtypes are transferable across independent datasets recorded under different conditions. The subtype whose functional connectivity differed most from those of healthy controls was less responsive to psychotherapy treatment for PTSD and failed to respond to antidepressant medication for MDD. By contrast, both subtypes responded equally well to two different forms of repetitive transcranial magnetic stimulation therapy for MDD. Our data-driven approach may constitute a generalizable solution for connectome-based diagnosis. |
|
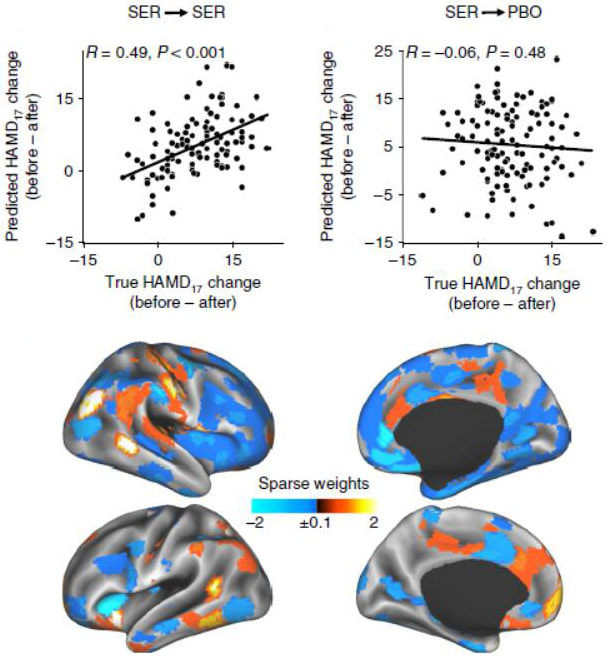
Wei Wu, Yu Zhang, Jing Jiang, Molly Lucas, Gregory Fonzon, Camarin Rolle, Crystal Cooper, Cherise Chin-Fatt, Noralie Krepel, Carena Cornelssen, Rachael Wright, Russel Toll, Hersh Trivedi, Karen Monuszko, Trevor Caudle, Kamron Sarhadi, Manish Jha, Joseph Trombello, Thilo Deckersbach, Phil Adams, Patrick McGrath, Myrna Weissman, Maurizio Fava, Diego Pizzagalli, Martjin Arns, Madhukar Trivedi, Amit Etkin
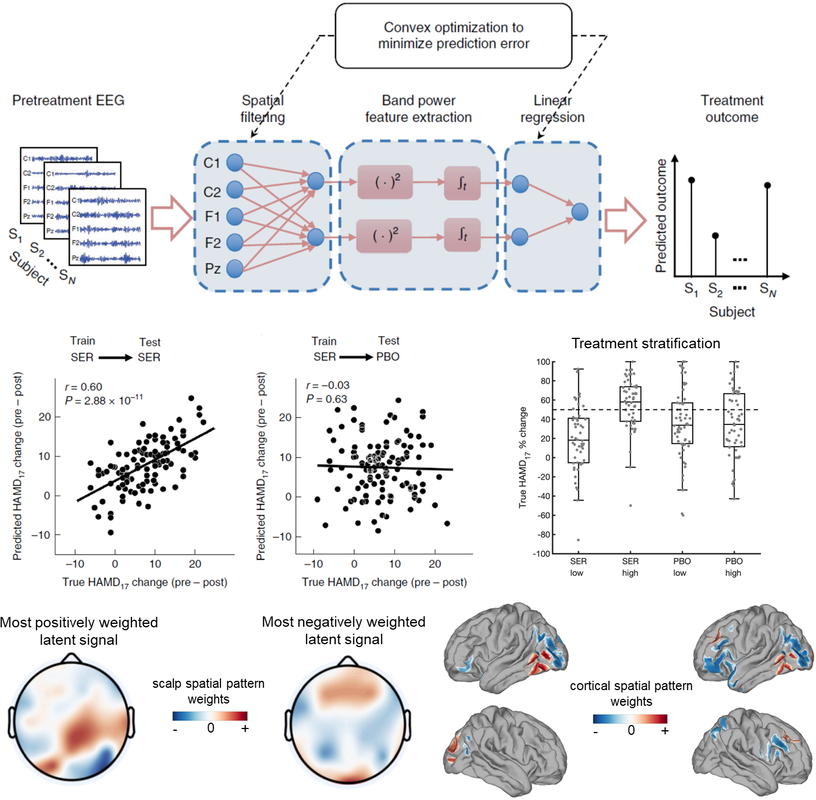
Nature Biotechnology, 2020 Problem: Antidepressants are widely prescribed, but their efficacy relative to placebo is modest, in part because the clinical diagnosis of major depression encompasses biologically heterogeneous conditions. Solution: We designed a latent-space machine-learning algorithm tailored for resting-state EEG and applied it to data from the largest imaging-coupled, placebocontrolled antidepressant study. Results: Our computational model robustly predicted treatment outcome with the antidepressant sertraline and distinguishing between response to sertraline versus placebo at the individual patient level and which may furthermore support treatment selection between medication and rTMS. Together, these findings ground in individual-level neurobiology a treatment-responsive phenotype obscured within the broader clinical diagnosis of depression and its associated biological heterogeneity and lay a path toward machine-learning-driven personalized approaches to treatment in depression. |
|
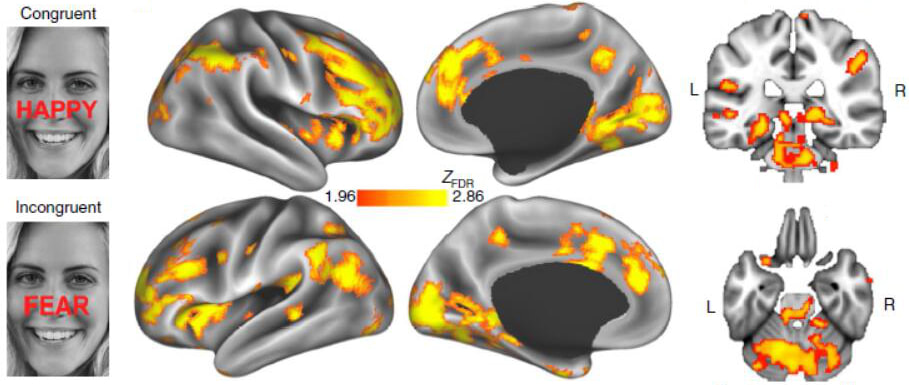
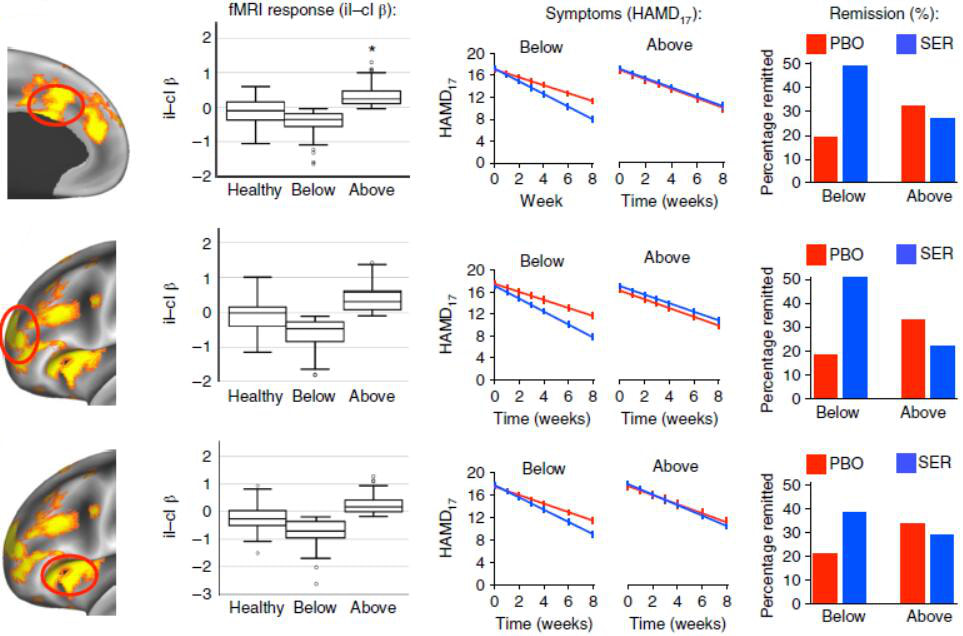
Gregory Fonzon#, Amit Etkin#, Yu Zhang#, Wei Wu, Crystal Cooper, Cherise Chin-Fatt, Manish Jha, Joseph Trombello, Thilo Deckersbach, Phil Adams, Melvin Mclnnis, Patrick McGrath, Myrna Weissman, Maurizio Fava, Maduhkar Trivedi
Nature Human Behaviour, 2019 (#Co-first authors) Problem: The efficacy of antidepressant treatment for depression is controversial due to the only modest superiority demonstrated over placebo. However, neurobiological heterogeneity within depression may limit overall antidepressant efficacy. Solution: We propose to identify a neurobiological phenotype responsive to antidepressant treatment by testing pretreatment brain activation during response to, and regulation of, emotional conflict as a moderator of the clinical benefit of the antidepressant sertraline versus placebo. Results: Using neuroimaging data from a large randomized controlled trial, we found widespread moderation of clinical benefits by brain activity during regulation of emotional conflict, in which greater downregulation of conflict-responsive regions predicted better sertraline outcomes. Treatment-predictive machine learning using brain metrics outperformed a model trained on clinical and demographic variables. |